Documentation
Common Data Model (CDM)
Convert Database to CDM (ETL)
Tool Specific Documentation
Common Data Model (CDM)
Convert Database to CDM (ETL)
Tool Specific Documentation
NOTICE FEB 2020: the current documentation can be found on Github http://ohdsi.github.io/WhiteRabbit
WhiteRabbit is a software tool to help prepare for ETLs (Extraction, Transformation, Loading) of longitudinal healthcare databases into the Observational Medical Outcomes Partnership (OMOP) Common Data Model (CDM). The source data can be in comma-separated text files, or in a database (MySQL, SQL Server, Oracle, PostgreSQL, Microsoft APS, Microsoft Access, Amazon RedShift, Google BigQuery). Note that the CDM will need to be in one of a limited set of database platforms (SQL Server, Oracle, PostgreSQL, Microsoft APS, Amazon RedShift, Google BigQuery).
WhiteRabbit’s main function is to perform a scan of the source data, providing detailed information on the tables, fields, and values that appear in a field. This scan will generate a report that can be used as a reference when designing the ETL, for instance by using the Rabbit-In-a-Hat tool. White Rabbit differs from standard data profiling tools in that it attempts to prevent the display of personally identifiable information (PII) data values in the generated output data file.
The typical sequence for using this software to scan source data in preparation of developing an ETL into an OMOP CDM:
Once the scan report is created, this report can then be used in the Rabbit-In-a-Hat tool or as a stand-alone data profiling document.
All source code and installation instructions available on GitHub: https://github.com/OHDSI/WhiteRabbit
Any bugs/issues/enhancements should be posted to the GitHub repository: https://github.com/OHDSI/WhiteRabbit/issues
Any questions/comments/feedback/discussion can be posted on the OHDSI Developer Forum: http://forums.ohdsi.org/c/developers
Any files that WhiteRabbit creates will be exported to this local folder. Use the “Pick Folder” button to navigate in your local environment where you would like the scan document to go.
Here you can specify the location of the source data. The following source types are supported: delimited text files, MySQL, Oracle, SQL Server, PostgreSQL, and Google BigQuery. Below are connection instructions for each data type of data source. Once you have entered the necessary information, the “Test connection” button can ensure a connection can be made.
Delimited text files
WhiteRabbit will look for the files to scan in the same folder you set up as a working directory.
MySQL
Oracle
SQL Server
When the SQL Server JDBC drivers are installed, you can also use Windows authentication. In this case, user name and password should be empty.
MS APS
At this time WhiteRabbit does not run on this platform.
PostgreSQL
Google BigQuery
Google BigQuery (GBQ) supports two different connection/authentication methods – application default credentials and service account authentication. The former method is considered more secure because it writes auditing events to stackdriver. The specific method used is determined by the arguments provided to the configuration panel as described below.
Authentication via application default credentials:
When using application default credentials authentication, you must run the following gcloud command in the user account only once: 'gcloud auth application-default login' (do not include the single quote characters). An application key is written to ~/.config/gcloud/application_default_credentails.json.
Authentication via service account credentials:
A scan generates a report containing information on the source data that can be used to help design the ETL. Using the Scan tab in WhiteRabbit you can either select individual tables in the selected source database by clicking on ‘Add’ (Ctrl + mouse click), or automatically select all tables in the database by clicking on ‘Add all in DB’.
There are a few setting options as well with the scan:
Once all settings are completed, press the “Scan tables” button. After the scan is completed the report will be written to the working folder.
For various reasons one could prefer to run WhiteRabbit from the command line. This is possible by specifying all the options one would normally select in the user interface in an .ini file. An example ini file can be found here. Then, we can reference the ini file when calling WhiteRabbit from the command line:
java -jar WhiteRabbit.jar -ini WhiteRabbit.ini
After the scan is completed, a “ScanReport” Excel document will be created in the working folder location selected earlier. The document will have multiple tabs, one as an “Overview” and then one tab for each database table or delimited text files selected for the scan. The “Overview” tab will tell you about each table selected, what the columns in each table are, the data type of the columns, the amount of data within the table, the number of rows scanned, and the fraction of data empty. Below is an example image of the “Overview” tab. Column A will list what table the information is about, Column B the column being reviewed, Column C, the type of the column, Column E is the number of rows (however with text files it will return - 1), Column F will tell you how many rows of the N rows were reviewed, and Column G will let you know how many rows are empty.
For a tab that describes a single table, the columns names from the source table (or delimited text file) will be across the columns of the Excel tab. Each source table column will generate two columns in the Excel. One column will list all distinct values that have a “Min cell count” greater than what was set at time of the scan (Section - Performing the Scan). If a list of unique values was truncated, the last value in the list will be “List truncated”; this indicates that there are one or more additional unique source values that appear less than the number entered in the “Min cell count” described in Section - Performing the Scan. Next to each distinct value will be a second column that contains the frequency, or the number of times that value occurs in the data. These two columns (distinct values and frequency) will repeat for all the source columns in the table profiled in the workbook.
The report is powerful in understanding your source data by highlighting what exists. For example, if the following results were given back on the “SEX” column within one of the tables scanned, we can see that there were two common values (1 and 2) that appeared 61,491 and 35,401 times respectively. WhiteRabbit will not define 1 as male and 2 as female; the data holder will typically need to define source codes unique to the source system. However these two values (1 & 2) are not the only values present in the data because we see this list was truncated. These other values appear with very low frequency (defined by “Min cell count”) and often represent incorrect or highly suspicious values. When generating an ETL we should not only plan to handle the high-frequency gender concepts 1 and 2 but the other low-frequency values that exist within this column.
This feature allows one to create a fake dataset based on a WhiteRabbit scan report. This dataset could be used to develop ETL code when direct access to the data is not available.
This feature is currently still experimental.
Rabbit-In-a-Hat comes with WhiteRabbit and is designed to read and display a WhiteRabbit scan document. WhiteRabbit generates information about the source data while Rabbit-In-a-Hat uses that information and through a graphical user interface to allow a user to connect source data to tables and columns within the CDM. Rabbit-In-a-Hat generates documentation for the ETL process it does not generate code to create an ETL.
The typical sequence for using this software to generate documentation of an ETL:
Rabbit-In-a-Hat comes with WhiteRabbit, refer to WhiteRabbit's installation section.
To create a new document, navigate to File –> Open Scan Report. Use the “Open” window to browse for the scan document created by WhiteRabbit. When a scan document is open, the tables scanned will appear in orange boxes on the “Source” side of the Tables.
Save the Rabbit-In-a-Hat document by going File –> Save as.
To open an existing Rabbit-In-a-Hat document use File –> Open ETL specs.
Rabbit-In-a-Hat allows you to select which CDM version (v4 or v5) you'd like to built your ETL specification against.
See the graphic below for how to select your desired CDM:
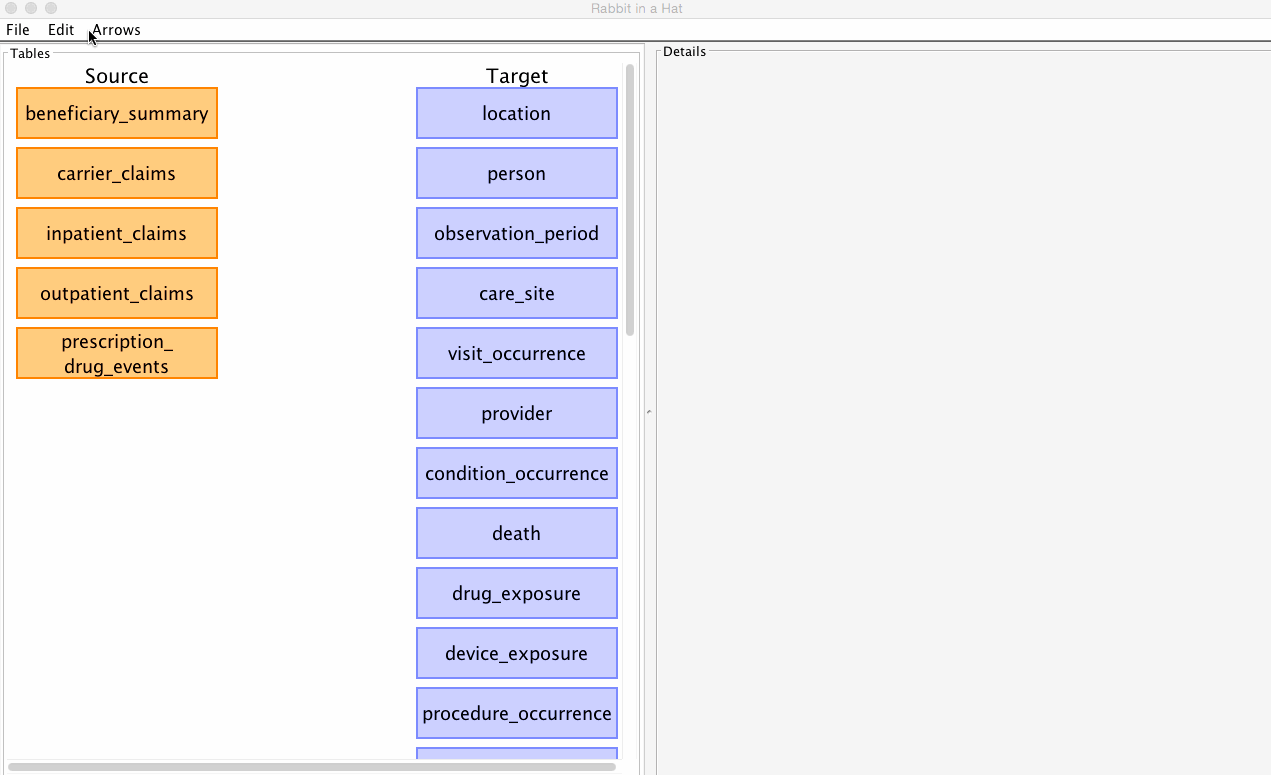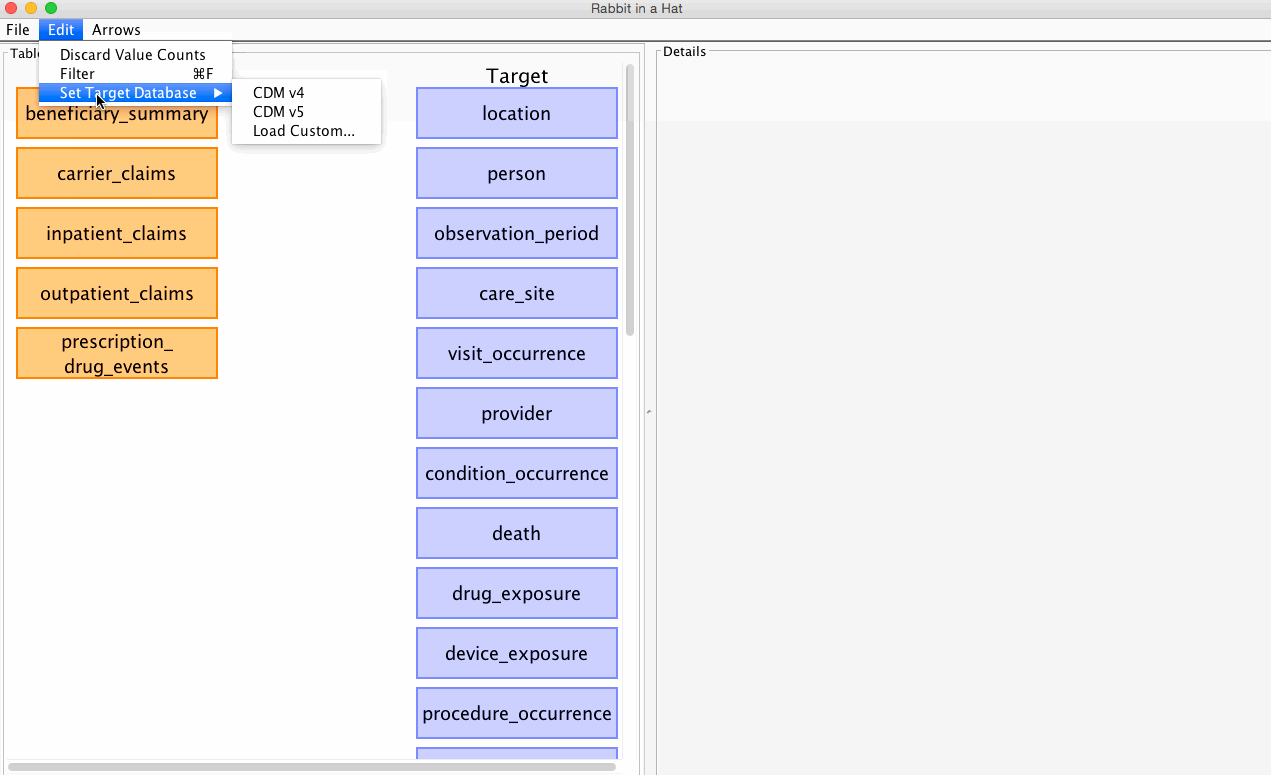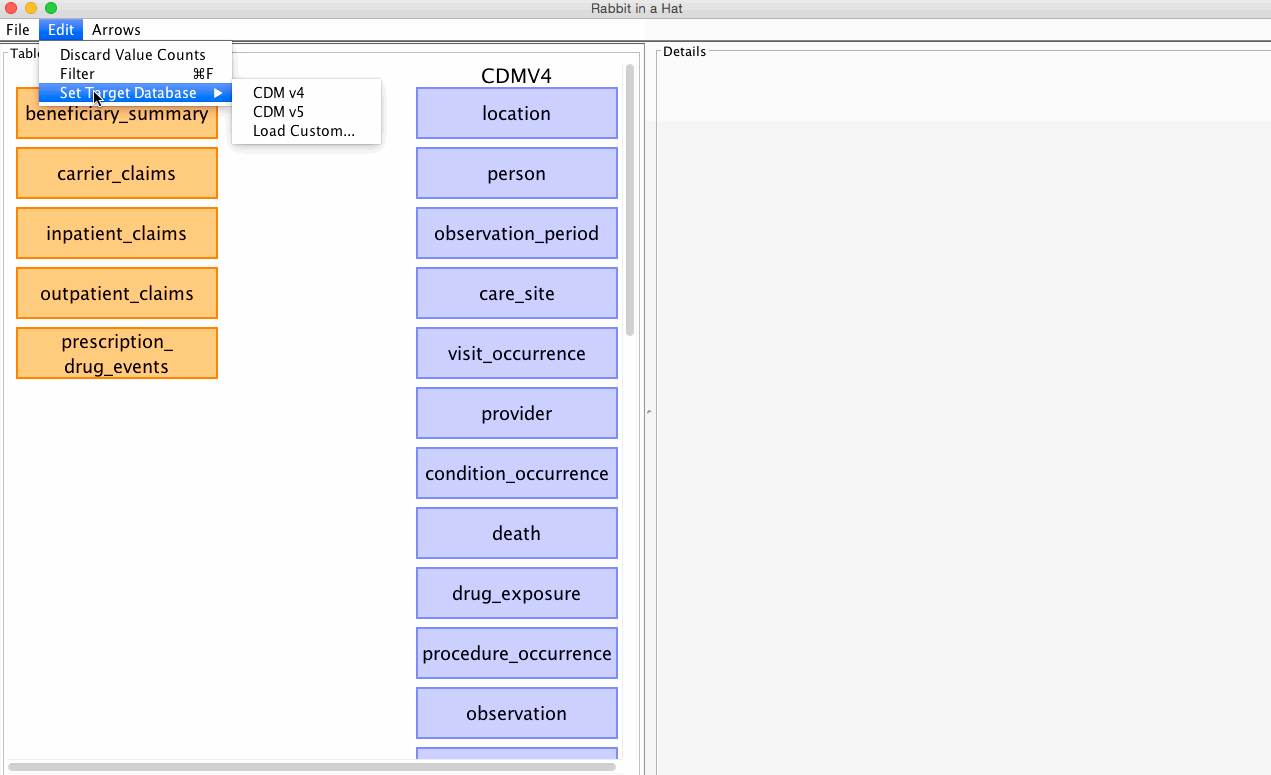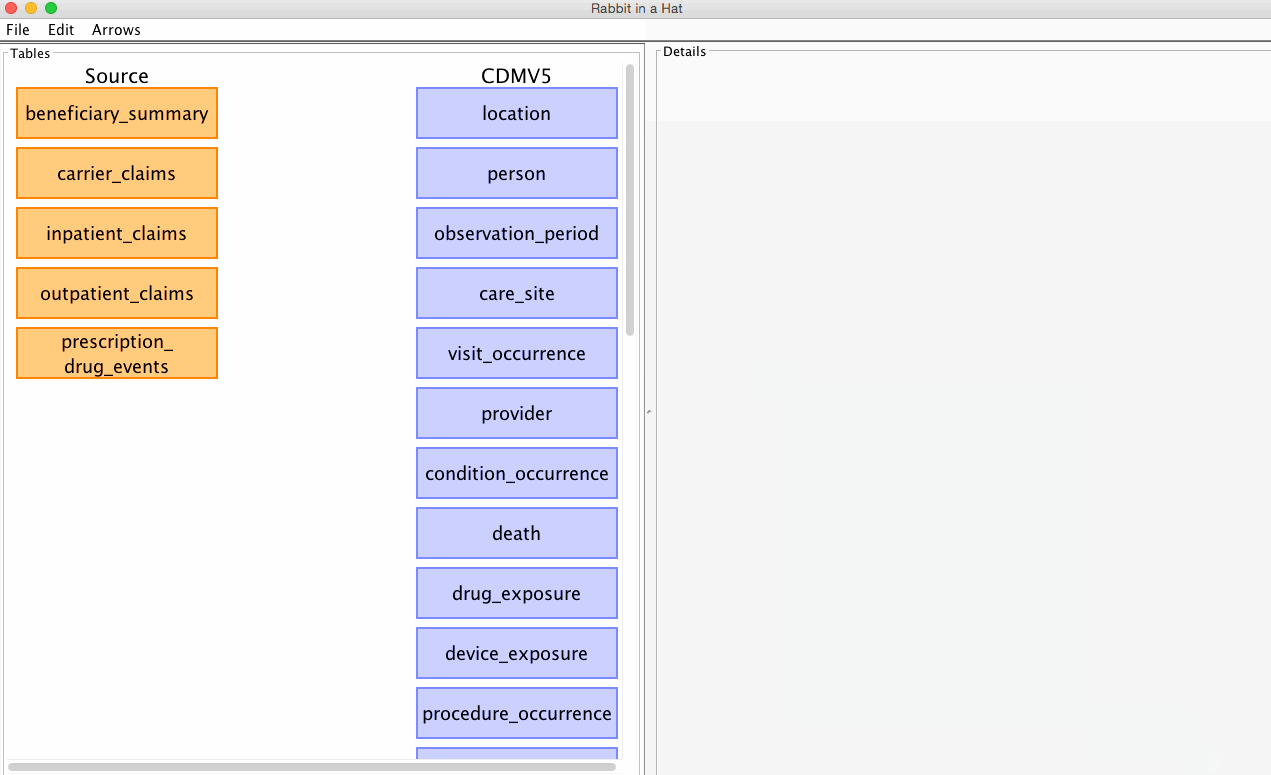
The CDM version can be changed at any time, but beware that some of your existing mappings may be lost in the process. By default, Rabbit-In-a-Hat will attempt to pereserve as many mappings between the source data and the newly selected CDM as possible. When a new CDM is selected, Rabbit-In-a-Hat will drop any mappings if the mapping's CDM table or CDM column name no longer exist
For instance, switching from CDMv4 to CDMv5, a mapping from source to CDM person.person_source_value will be kept because the person table has person_source_value in both CDMv4 and CDMv5. However, person.assocaited_provider_id exists only in CDMv4 and has been renamed to person.provider_id in CDMv5 and so that mapping will not be kept when switching between these two CDMs.
There are times when users might need to load in a customized version of the CDM, for instance if they are sandboxing new features. To load in a custom CDM schema, first you must create a CSV file that uses the same format as the existing CDMv5 schema file.
Once you have created the CSV file, load it into RiaH as shown below:

Please note that the name of the file you load in becomes the label that appears above the target tables, so “My Super File.csv” will create the label “My Super File” above the target tables, so name your CSV accordingly.
It is assumed that the owners of the source data should be able to provide detail of what the data table contains, Rabbit-In-a-Hat will describe the columns within the table but will not provide the context a data owner should provide. For the CDM tables, if more information is needed navigate to the OMOP CDM web page (http://omop.org/CDM) and review the current OMOP specification.
To connect a source table to a CDM table, simply hover over the source table until an arrow head appears.
Use your mouse to grab the arrow head and drag it to the corresponding CDM table. In the example below, the drug_claims data will provide information for the drug_exposure table.
If you click on the arrow once it will highlight and a Details window will appear in the right pane. Use this to describe Logic or Comments that someone developing the ETL code should know about this source table to CDM table mapping.
Continue this process until all tables that are needed to build a CDM are mapped to their corresponding CDM tables. One source table can map to multiple CDM tables and one CDM table can receive multiple mappings. There may be tables in the source data that should not be map into the CDM and there may be tables in the CDM that cannot be populated from the source data.
By double clicking on an arrow connecting a source and CDM table, it will open a Fields pane below the arrow selected. The Fields pane will have all the source table and CDM fields and is meant to make the specific column mappings between tables. Hovering over a source table will generate an arrow head that can then be selected and dragged to its corresponding CDM field. For example, in the drug_claims to drug_exposure table mapping example, the source data owners know that patient_id is the patient identifier and corresponds to the CDM.person_id. Also, just as before, the arrow can be selected and Logic and Comments can be added.
If you select the source table orange box, Rabbit-In-a-Hat will expose values the source data has for that table. This is meant to help in the process in understanding the source data and what logic may be required to handle the data in the ETL. In the example below ndcnum is selected and raw NDC codes are displayed starting with most frequent (note that in the WhiteRabbit scan a “Min cell count” could have been selected and values below that frequency will not show).
Continue this process until all source columns necessary in all mapped tables have been mapped to the corresponding CDM column. Not all columns must be mapped into a CDM column and not all CDM columns require a mapping. One source column may supply information to multiple CDM columns and one CDM column can receive information from multiple columns.
To generate an ETL MS Word document use File –> Generate ETL document and select a location to save. It may take a moment before the document is finished creating. This document will contain all notes from Rabbit-In-a-Hat.
Once the information is in the document, if an update is needed you must either update the information in Rabbit-In-a-Hat and regenerate the document or update the document. If you make changes in the document, Rabbit-In-a-Hat will not read those changes and update the information in the tool. However it is common to generate the document with the core mapping information and fill in more detail within the document.
Once the document is completed, this should be shared with the individuals who plan to implement the code to execute the ETL.
To make sure the ETL process is working as specified, it is highly recommended to create unit tests that evaluate the behavior of the ETL process. To efficiently create a set of unit tests Rabbit-in-a-Hat can generate a testing framework.
The following lists best practices in using WhiteRabbit and Rabbit-In-a-Hat to manage your ETL documentation process: